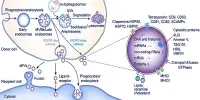
Ribosomes are the building blocks of life. Ribosomes are required by every cell on the planet to translate genetic data into the critical proteins required for the organism’s operation and, consequently, for the manufacture of new ribosomes. However, scientists are still unsure how these critical nanomachines are put together.
New high-resolution photos of the big ribosomal subunit give light on how nature’s most basic molecule coalesces in human cells. The discoveries, which were published in Science, bring us one step closer to understanding ribosome assembly.
“We now have a pretty good idea of how the large ribosomal subunit is assembled in humans,” explains Sebastian Klinge of Rockefeller. “We still have a lot of gaps in our understanding, but we have a much better idea now than we did before.”
Solving the large subunit
Ribosomes were discovered about 70 years ago at Rockefeller University. Researchers discovered that they are made up of two separate components: a small 40S subunit that interprets messenger RNA and a bigger 60S subunit that connects protein fragments. However, those were the broadest brushstrokes. The specific steps involved in assembling these complicated molecules into their mature state have long been a mystery.
Klinge’s long-term approach to this broader problem has been to figure out how ribosomes develop in the first place. Klinge’s lab was among the first to use cryo-electron microscopy to capture footage of a nonbacterial ribosome assembling towards its final shape, and the lab has since taken an even more granular approach—painfully stringing snapshots of maturing ribosomes together to understand how these molecules get from one point in their assembly to the next.
Klinge and colleagues have found and characterized over 200 ribosome assembly variables that govern ribosome modification, processing, and folding in recent years.
Klinge and colleagues focused on the human big ribosomal subunit (60S) for the current investigation. The team already knew from yeast studies that the large subunit is formed by two precursors (a 5S rRNA and a 32S pre-rRNA snapping together), but “we wanted to know all of the events that need to happen for this to occur,” says Arnaud Vanden Broeck, a postdoctoral researcher in Klinge’s lab. “We wanted to explain how the large subunit is assembled and processed in human cells.”
Vanden Broeck and Klinge used novel genome editing and biochemistry approaches to acquire high-resolution cryo-EM structures of 24 human large ribosomal subunit assembly intermediates as they matured. The graphics depict how assembly factors, proteins, and enzymes interact with RNA components to induce 60S creation and maturation. The findings provide a nearly full description of how the human large subunit is assembled.
“For sixty years, we had almost nothing on the intermediates that form the human 60S—it was all but invisible to us—and now we’ve jumped from nothing to pretty good coverage,” Vanden Broeck says, admitting that some of the rarest and most transient steps on the road to the mature 60S may have eluded the team and fallen through the cracks. “We still have a lot of work to do.”
Nonetheless, major discoveries from the study may already be influencing related fields of research. Among the found intermediary steps are signaling pathways that suggest a relationship between ribosome assembly and cellular metabolism, implying that a thorough knowledge of ribosomes may require close collaboration with cell metabolism experts. Furthermore, the study’s granular look into the processes of ribosome production may provide a vital background for scientists studying disorders connected to ribosome mutations.
For the time being, Klinge and Vanden Broeck are pleased to marvel at the significant progress. “It’s not guesswork anymore,” Klinge explains. “We can now see in great detail what happens when the large subunit assembles.” It’s humbling to understand that we can finally see what generates ribosomes and drives protein production in all of our cells.”