Scientists describe a method for stable attachment of peptides to tRNAs, which has allowed them to gain new fundamental insights into ribosome function by determining the atomic-level structures of ribosomes and the shapes that peptides take inside the ribosome.
Chains of genetic material called messenger RNAs (mRNAs) are matched with the corresponding transfer RNAs (tRNAs) inside tiny cellular machines called ribosomes to create amino acid sequences that exit the ribosome as proteins. Unfinished proteins are known as nascent chains, and they are left attached to the ribosome.
Scientists know that some of these nascent chains can regulate ribosome activity and that they can sometimes interfere with antibiotics, many of which work by targeting bacterial ribosome activity. Scientists are unsure why this occurs, primarily because it is difficult to visualize the ribosome-peptide-drug interactions while the unfinished proteins are still attached to the ribosome.
Scientists at the University of Illinois Chicago are the first to report a method for stable attachment of peptides to tRNAs, which has allowed them to gain new fundamental insights into ribosome function by determining the atomic-level structures of ribosomes and the shapes that these peptides take inside the ribosome.
Their method is newly reported in the journal Nature Chemistry.
The method has been used for a long time in chemistry, but it has never been applied in this way. It’s mimicking nature, basically, and with our advanced imaging experience, we are now seeing how nature works at a high resolution.
Yury Polikanov
“The challenge has been to see up close the structure of the ribosome and the exit tunnel in the presence of the nascent peptides because, in nature, the ribosome is very quick for us to capture images or conduct experiments,” said Yury Polikanov, associate professor in the biological sciences department at the College of Liberal Arts and Sciences. “Until the advent of this new method, we’ve essentially been blinded from seeing what is happening in the active site of the ribosome at this critical moment in time.”
Polikanov and his colleague Egor Syroegin, a PhD candidate in biological sciences at UIC, used a technique known as native chemical ligation to fuse custom peptides with the tRNA, resulting in a peptidyl-tRNA.
“Obtaining tRNA molecules linked to peptides, similar to those found inside the ribosome during protein synthesis, has been a long-held dream of many researchers in the field for nearly two decades,” Polikanov said. “This has been extremely difficult because no enzymes can directly attach peptides to tRNA.”
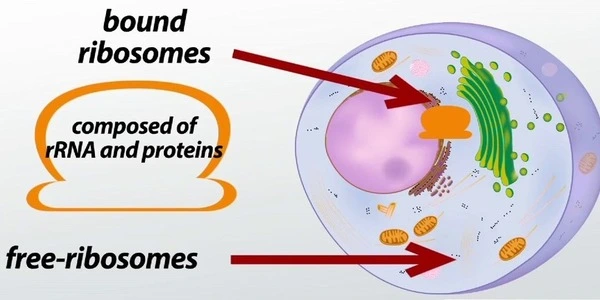
“The method has been used for a long time in chemistry, but it has never been applied in this way. It’s mimicking nature, basically, and with our advanced imaging experience, we are now seeing how nature works at a high resolution,” Syroegin said.
With this new approach, Polikanov and Syroegin determined a set of high-resolution structures of the ribosome carrying peptidyl-tRNAs of various lengths.
According to Polikanov, detailed analysis of these structures provides new and surprising insights into the mechanism of the ribosome’s catalytic center and answers several long-standing fundamental questions in the ribosome field.
“We saw that depending on the sequence, different peptides can form different shapes or folds within the ribosomal tunnel, and we can synthesize different peptides of different sequences and then follow their shape very precisely, because of the high resolution of our structures,” Syroegin explained. “So we can confidently state that ‘these peptides in this sequence have this shape’ or ‘another peptide has a different shape.’ This is significant because nascent peptide folding determines whether drugs will arrest ribosomes or not.”
“This method opens up a plethora of opportunities for structural and functional studies aimed at understanding the mechanisms of ribosome functioning, as well as sequence-specific ribosome stalling caused by certain antibiotics,” Polikanov said.
The paper, “Insights into ribosome function from the structures of non-arrested ribosome nascent chain complexes,” was co-authored by Polikanov and Syroegin, as well as Elena Aleksandrova, a research specialist in the biological sciences department at UIC.