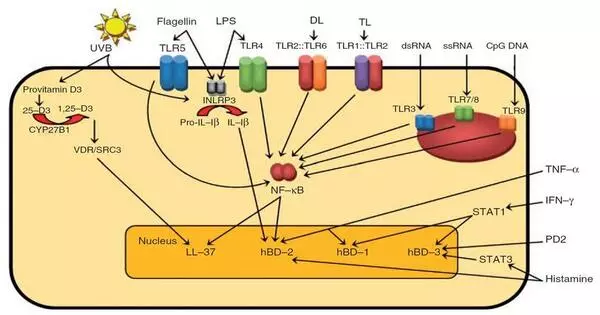
Antimicrobial peptides (AMPs) are a class of small peptides found throughout nature that play an important role in the innate immune systems of various organisms. AMPs have a wide range of inhibitory effects on bacteria, fungi, parasites, and viruses.
Researchers at Princeton University’s Department of Chemistry have discovered a new multi-step pathway by which bacteria in the mammalian gut produce antimicrobial peptides. The newly discovered biosynthetic pathway converts a biologically inert peptide into structurally complex antibiotics known as enteropeptins. Enteropeptins are a type of ribosomally synthesized peptide natural product known as a RiPP.
The ribosome, which is limited to the 20 canonical amino acids, synthesizes the core structure of these products. Within enteropeptin, the Mo Lab discovered and characterized new metalloenzymes capable of converting arginine, a canonical amino acid, into N-methylornithine, a noncanonical amino acid.
This is the first time a RiPP natural product containing this unusual amino acid has been reported. Professor Mohammad Seyedsayamdost’s lab made the discovery. “Biosynthesis-guided discovery reveals enteropeptins as alternative sactipeptides containing N-methylornithine,” the lab’s paper, was published in Nature Chemistry last month.
It’s a new approach to natural product discovery in that you start with the biosynthetic gene cluster and then do in-depth analysis of each enzyme’s reaction to understand all the different transformations that are going on.
Covington Clark
Kenzie Clark, the paper’s first author and a former graduate student at the Mo Lab, explained the key findings.
“The ribosome generates a precursor peptide, which is then acted on by metalloenzymes encoded in the same gene cluster,” Clark explained. “A series of metalloenzymes — in this case, three of them — convert arginine into N-methylornithine step by step to produce enteropeptin.” The interesting thing is that the peptide has no biological activity on its own. However, once these modifications are made, it becomes this active biomolecule that effectively inhibits the growth of the producing strain.”
Brett Covington, a postdoctoral researcher in the lab and co-author on the paper, weighed in on this finding.
“That’s a trend we are seeing with a lot of these RiPP products that we’ve uncovered: they have very narrow-spectrum activity and they tend to inhibit the growth of the organism that’s making the compound,” he said. “That was the case here with enteropeptin. It really only inhibited the Enterococcus that made enteropeptin. Why these Enterococci make an antibiotic peptide to inhibit their own growth is a question that we’re pursuing. They are likely involved in generating a more recalcitrant bacterial population.”
Working Through Gene Clusters
The study fits right in with the Mo Lab’s mission of discovering new bacterial natural products and understanding how they are biosynthesized. In 2018, the Mo Lab published a paper in the Journal of the American Chemical Society (JACS) in which they used bioinformatics to identify 600 RiPP gene clusters from streptococci that use radical S-adenosylmethionine (rSAM) enzymes, one of nature’s largest-known families of enzymes. While it is a large group with over 500,000 members and is found in all three kingdoms of life, most of these enzymes are not well understood.
Based on similar precursor peptide sequences, the researchers classified 600 newly discovered RiPP gene clusters into 16 subfamilies. In the lab, they began working through these families, discovering new reactions and interesting chemistry along the way.
“It’s a new approach to natural product discovery in that you start with the biosynthetic gene cluster and then do in-depth analysis of each enzyme’s reaction to understand all the different transformations that are going on,” Covington explained. “Then we look for the mature product in the bacterial host, which includes all of these different enzymatic transformations. It’s a novel approach.”
So far, they’ve only worked with clusters that code for a single metalloenzyme. This latest work targets multi-step pathways with more complex chemistries that have not just one interesting reaction, but three catalzyed by three different classes of metalloenzymes. Two are iron-dependent and one is manganese-dependent.
“One of these metalloenymes generates a very reactive intermediate that’s able to do really challenging chemical reactions, for example carbon-carbon bond formation at unactivated carbon centers,” said Clark.
“Many of the structures they generate would be extremely difficult for synthetic chemists to create. But the third enzyme in the pathway was also intriguing. It was not classified as a known enzyme family. It was discovered that a four-iron-four-sulfur cluster and cofactor SAM were used to N-methylate ornithine, a novel and very interesting modification reaction.”
Covington continued: “One of the things I like best about working here at the Mo lab is the rush you get when you finally find one of these products. All of the work is done on the bench to understand the reaction in the test tube, and it’s extremely satisfying to see it match up with what’s going on inside the bacteria.”