Neuroscientists can better identify brain cells by using existing tools such as graphical models. The algorithm has significant implications for developmental diseases such as Alzheimer’s because scientists can find interventions once they understand the mechanism of a disease. The algorithm greatly accelerates the rate at which whole-brain data can be analyzed and supports crowdsourcing, allowing the larger neuroscience community to test the algorithm and build atlases.
Neuroscientists frequently struggle to accurately identify cells needed to understand brain activity that causes behavior changes such as declining memory or impaired balance and tremors when researching the causes and potential treatments for degenerative conditions such as Alzheimer’s or Parkinson’s disease.
Using existing tools such as graphical models, a multidisciplinary team of Georgia Institute of Technology neuroscience researchers discovered a better way to identify cells and understand disease mechanisms, potentially leading to better understanding, diagnosis, and treatment. Their findings were published in the journal eLife. The National Institutes of Health and the National Science Foundation funded the study.
Neuroscientists identifying cells to better understand healthy and diseased behavior. The field of neuroscience studies how the nervous system functions, and how genes and environment influence behavior.
Neuroscience is the study of how the nervous system works and how genes and environment influence behavior. Neuroscientists hope to find cures for diseases by using new technologies to understand the natural and dysfunctional states of biological systems. Before that can happen, neuroscientists must first understand which brain cells are driving behavior, but mapping brain activity cell by cell isn’t as easy as it appears.
No Two Brain Cells Are Alike
Scientists have traditionally established a coordinate system to map each cell location by comparing images to an atlas, but the notion in literature that “all brains look the same is absolutely not true,” according to Hang Lu, the Love Family Professor of Chemical and Biomolecular Engineering at Georgia Tech.
Taking a coordinate approach presents two main challenges: first, the sheer number of cells in which none look that distinct; second, cells vary from individual to individual. “This is a current huge bottleneck – you can record neuron activities all you want but if you don’t understand which cells are doing what, it’s difficult to compare between brains or conditions and draw meaningful conclusions,” Lu said.
There are also noises in data that make establishing correspondence between two different regions of the brain difficult, according to graduate researcher Shivesh Chaudhary. “Some deformations in the data may exist, or some portions of the shape may be missing,” he explained.

Concentrating on Cell Relationships Rather Than Geography
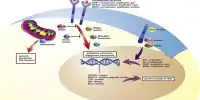
To overcome these obstacles, the Georgia Tech researchers combined two disciplines – graphical models in machine learning and metric geometry approaches to shape matching in mathematics – to develop a computational method for identifying cells in their model organism, the nematode C.elegans.
To create their own modeling software, the team used frameworks from other fields such as natural language processing. The computer can determine what sentences mean in natural language processing by capturing dependencies between words in a statement.
The researchers used a similar model, but instead of capturing dependencies between words, “we captured them among the neurons to identify cells,” Chaudhary explained, noting that this approach limits error propagation when compared to other methods that examine each cell’s geographic location. “Using cell relationships was actually more useful in defining a cell’s identity,” Lu said. “Defining one has implications for the identities of the other cells.”
According to the research team, the approach is far more accurate than the current method of identification. While not perfect, the algorithm performs significantly better in the face of imperfect data and “gets less rattled” by noise or errors, according to Lu.
The algorithm has far-reaching implications for many developmental diseases, because once scientists understand the mechanism of a disease, they can develop treatments. “This can be used to perform drug and genetic screenings to assess genetic risks. You can examine how someone’s genetic background causes cells to behave differently than the standard reference genetic background “Lu stated.
“One interesting aspect of this approach is that it is data-driven, so it captures the differences between individual worms. This method has a high potential for use in a variety of studies on development and function under normal and disease-like conditions “Yun Zhang, professor in the Department of Organismic and Evolutionary Biology at Harvard University’s Center for Brain Science, agreed.
Faster Data Analysis
The algorithm significantly increases the speed with which whole-brain data can be analyzed. Prior to this advancement, the researchers explained that their lab might take 20 minutes to record a set of data, but it would take weeks to identify cells and analyze data. The algorithm takes “at most overnight on a desktop,” according to Chaudhary.
The technique also supports crowdsourcing, collaborative online platforms that expose the algorithm to a larger community, allowing it to be tested and atlases built. “Every researcher working on the same problem could make recordings and contribute to the further development of these atlases, which will be widely usable in all contexts,” Lu explained.
The researchers attribute the project’s success to their ability to draw on multiple disciplines, including physics, biology, math, and chemistry. Chaudhary, who has a bachelor’s degree in chemical engineering, used advances in computer science and math to solve this particular neuroscience problem.
“We have a physicist working on microscopes, biologists, and people like me who are more interested in computer science in our labs. We also work with a pure mathematician “He elaborated. “Everything is available in the field of neuroscience. You can go in any direction you want.”