Children’s Hospital of Philadelphia (CHOP) researchers have created a versatile and low-cost technology for targeted sequencing of full-length RNA molecules, which could speed up the discovery of new diagnostics and therapies. The technology, known as TEQUILA-seq, is far less expensive than currently available options for targeted RNA sequencing and may be customized for a variety of academic and therapeutic applications. The specifics are reported in a paper published in Nature Communications.
Before being translated into a protein, an RNA molecule might be cut and linked in a variety of ways on its path from gene to protein. This is known as alternative splicing, and it enables a single gene to encode many proteins. Alternative splicing occurs in many biological processes, but it can be dysregulated in disorders such as cancer, resulting in harmful RNA molecules. Researchers require a precise count of all the RNA molecules (known as “transcript isoforms”) that come from a particular gene to understand how alternative splicing can cause disease.
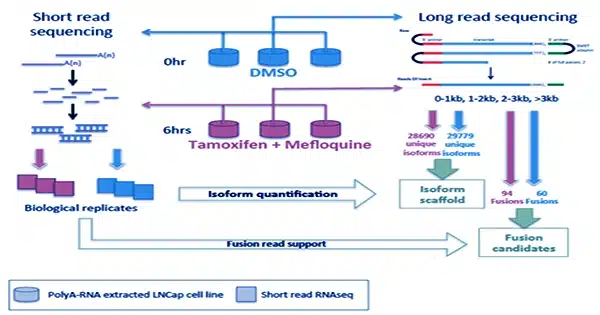
One method is to use “long-read” RNA sequencing systems, which sequence RNA molecules exceeding 10,000 bases long end-to-end, catching all transcript isoforms. However, the low sequencing yield of these long-read platforms has limited their general use, particularly in the clinical setting, where obtaining long-read RNA sequencing data at clinically meaningful depth could be prohibitively expensive. Targeted sequencing, which involves enriching certain nucleic acid sequences of interest prior to sequencing, is a valuable method that can significantly improve coverage of predefined targets; nonetheless, the expense and complexity of target capture have been impediments to wider adoption.
“Targeted long-read RNA sequencing is an effective method for elucidating the RNA repertoire of any predefined set of genes.” Existing technologies for targeted sequencing of full-length RNA molecules, on the other hand, are either expensive or difficult to set up, putting them out of reach for many labs,” said co-senior author Lan Lin, Ph.D., Assistant Professor of Pathology and Laboratory Medicine at CHOP and a member of the Raymond G. Perelman Center for Cellular and Molecular Therapeutics. “TEQUILA-seq addresses this issue by being both affordable and simple to use. Users can customize the technique for various applications, and researchers can choose which genes to sequence and create the reagents for target capture in their own facilities. This has the potential to speed up the development of new diagnostic and therapeutic treatments for a variety of ailments.”
Hybridization capture-based enrichment, which uses small bits of nucleic acid termed oligonucleotides as capture probes, is one method that allows for targeted sequencing. These oligonucleotides (also known as “oligos”) are biotin-tagged and intended to hybridize to their targets based on nucleic acid sequence complementarity, allowing enabling simple capture and isolation of target sequences from biological samples. However, while hybridization capture-based enrichment is an effective method for targeted sequencing, commercially produced biotinylated capture probes are costly and can only be utilized for a limited number of reactions, making the per-sample cost for each capture process prohibitive.
To overcome this constraint, the CHOP researchers created TEQUILA-seq (Transcript Enrichment and Quantification Using Isothermal Linear-Amplified Probes in Conjunction with Long-Read Sequencing). A fundamental innovation in TEQUILA-seq is a nicking-endonuclease-triggered isothermal strand displacement amplification reaction, which can generate vast amounts of biotinylated capture probes from a low-cost pool of non-biotinylated oligos as templates. The researchers can synthesize 25 ug of TEQUILA probes from only 2 ng of template oligos, which can be employed in at least 250 capture reactions. Because of this novel approach to synthesizing capture probes, TEQUILA-seq is both cost-effective and scalable for broad target panels and large biological samples.
To test its performance, the researchers ran TEQUILA-seq on synthetic RNAs or human RNAs for several gene panels. In target capture and enrichment, TEQUILA probes outperformed commercial capture probes while costing hundreds of times less per capture reaction. Furthermore, the researchers revealed that TEQUILA-seq can significantly improve detection while maintaining the quantification of target RNA molecules.
“Using cheap reagents and an easy experimental workflow, TEQUILA-seq allows us to deeply sequence the full-length RNA molecules for any gene set across many biological samples,” said co-senior author Yi Xing, Ph.D., head of CHOP’s Center for Computational and Genomic Medicine. “This is very exciting, and it enables a wide range of medical applications, from RNA-guided genetic diagnosis to therapy development.”
The researchers used TEQUILA-seq to profile full-length RNA molecules of 468 actionable cancer genes across 40 breast cancer cell lines to demonstrate its biological value. They uncovered newly unknown transcript isoforms in cancer genes that have been extensively researched, which may give light on how genes that protect the body from cancer are inactivated in specific tumors.
“Our findings demonstrate that TEQUILA-seq can be widely applied for targeted sequencing of full-length RNA molecules,” Dr. Lin remarked. “In addition, TEQUILA probes are multipurpose capture probes.” They work with both long-read and short-read sequencing technologies for targeted RNA and DNA sequencing. The capacity to quickly and inexpensively create vast quantities of biotinylated capture probes for any target panel at a cheap cost and with ease can promote large-scale and population-level studies for a variety of fundamental, translational, and clinical applications.”